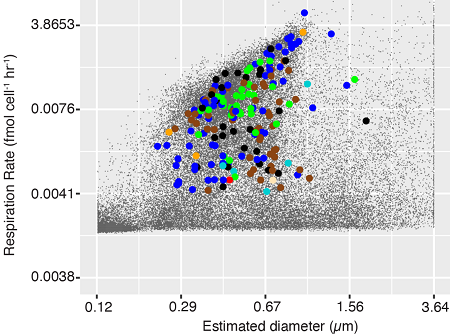
A study led by researchers from Bigelow Laboratory for Ocean Sciences, using SCGC’s new technology for integrated phenotype analyses and genomic sequencing of individual microbial cells, showed that cell-specific respiration rates differ by more than 1,000× among genera of marine prokaryoplankton. The majority of respiration was found to be performed by minority members of prokaryoplankton (including the Roseobacter cluster), whereas cells of the most prevalent lineages (including Pelagibacter and SAR86) had extremely low respiration rates. The decoupling of respiration rates from abundance among lineages, elevated counts of proteorhodopsin transcripts in Pelagibacter and SAR86 cells and elevated respiration of SAR86 at night indicate that proteorhodopsin-based phototrophy probably constitutes an important source of energy to prokaryoplankton and may increase growth efficiency. These findings suggest that the dependence of prokaryoplankton on respiration and remineralization of phytoplankton-derived organic carbon into CO2 for its energy demands and growth may be lower than commonly assumed and variable among lineages.
Read the whole article in Nature.