Updated November 24, 2025
To complete a SCGC service inquiry or request, start here. Examples of SCGC service deliverables and public data releases can be found here. To download this page in PDF format, click here.
Table 1. List of SCGC services. All prices are in US dollars (USD). A 15% discount is applied to orders exceeding $50,000. A 20% discount is applied to all services for Bigelow Laboratory’s in-house projects. Service fees do not include shipping costs.
| Service | Cat # | Unit | Unit price* |
| FACS-Based SAG Generation and Sequencing | |||
| FACS SAG Generation WGA-X. Reduced price | S-201 | 384-well plate | $1,900 |
| FACS SAG Generation WGA-Y. Reduced price | S-202 | 384-well plate | $3,700 |
| Integrated single cell respiration measurement | S-203 | 384-well plate | $900 |
| FACS SAG WGS 1.8 Billion Reads | S-211 | ≤384 SAGs | $24,000 |
| FACS SAG WGS 0.4 Billion Reads | S-212 | ≤384 SAGs | $19,000 |
| FACS SAG WGS 0.1 Billion Reads | S-213 | ≤384 SAGs | $15,000 |
| FACS SAG WGS Add-on 1.8 Billion Reads | S-221 | ≤384 SAGs | $14,000 |
| FACS SAG WGS Add-on 0.4 Billion Reads | S-222 | ≤384 SAGs | $9,000 |
| FACS SAG WGS Add-on 0.1 Billion Reads | S-223 | ≤384 SAGs | $6,000 |
| Semi-Permeable Capsule (SPC) Generation and Sequencing NEW! | |||
| SPC SAG Generation and Barcoding | S-311 | ≤ 10,000 SAGs | $15,000 |
| SPC SAG Illumina Library Production | S-312 | ≤ 2,500 SAGs | $2,000 |
| SPC SAG WGS, 1.8 Billion Reads | S-321 | ≤ 10,000 SAGs | $16,000 |
| SPC SAG WGS, 0.4 Billion Reads | S-322 | ≤ 10,000 SAGs | $11,000 |
| SPC SAG WGS, 0.1 Billion Reads | S-323 | ≤ 10,000 SAGs | $8,000 |
| Other Services | |||
| Biosafety Level 2 Handling | S-004 | 384-well plate | $600 |
| FACS SAG Re-Arraying | S-005 | 96-well plate | $800 |
| Sample Cryoprotectant glyTE | S-009 | 10 mL | $100 |
| Consultation | S-011 | 1 hour | $260 |
| Customized Services | S-020 | Custom | Request a quote |
OVERVIEW
Single-cell genomics unveils the genomic blueprints of the most fundamental units of life. It is a powerful approach to analyze biochemical properties, evolutionary histories and the biotechnological potential of uncultured microorganisms, which constitute over 99% of biological diversity on our planet. Single cell genomics consists of a series of integrated processes, starting with appropriate sample collection and preservation, followed by physical separation, lysis, and whole genome amplification of individual cells, then proceeding to DNA sequencing and sequence interpretation (1). These processes are incorporated in the comprehensive suite of services offered by SCGC (Table 1).
The SCGC is the world’s first research and service center with the primary focus on the single cell genomics of microorganisms (see About on the SCGC website). We analyze microorganisms from diverse microbiomes, biosafety level 2 organisms, and organisms from hard-to-process environments, such as soil and the deep biosphere. SCGC also processes individual cells of humans and other multicellular organisms. In most cases, SCGC can process samples when target cells or particles are in an aqueous solution at a concentration greater than 1,000 per mL, are less than 40 micrometers in diameter, and are cryopreserved and shipped in accordance with SCGC recommendations (see Preparation and Shipment on the SCGC website). Samples that have not been brought into aquatic suspension by SCGC customers may be analyzed as a customized service.
The processing of exceedingly small DNA quantities makes single cell genomics highly susceptible to DNA contamination and amplification biases. At SCGC, we have developed techniques to monitor and minimize methodological artifacts at every step of our workflow. Cell sorting and DNA amplification are performed in a cleanroom, and all consumables are decontaminated using in-house methods (2). Control wells on each microplate are used to detect potential DNA contamination. To prevent index switching during SAG sequencing, multiplexed libraries contain dual 10 bp barcodes. Genome de novo assemblies are evaluated by diverse QC tools. The entire workflow is assessed for contamination and assembly errors using microbial benchmark cultures with diverse genome complexity and G+C content, indicating no non-target and undefined bases and average frequencies of mis-assemblies, indels, and mismatches at <5 per 100 kbp (2).
SCGC SERVICES: FACS-BASED SAGs
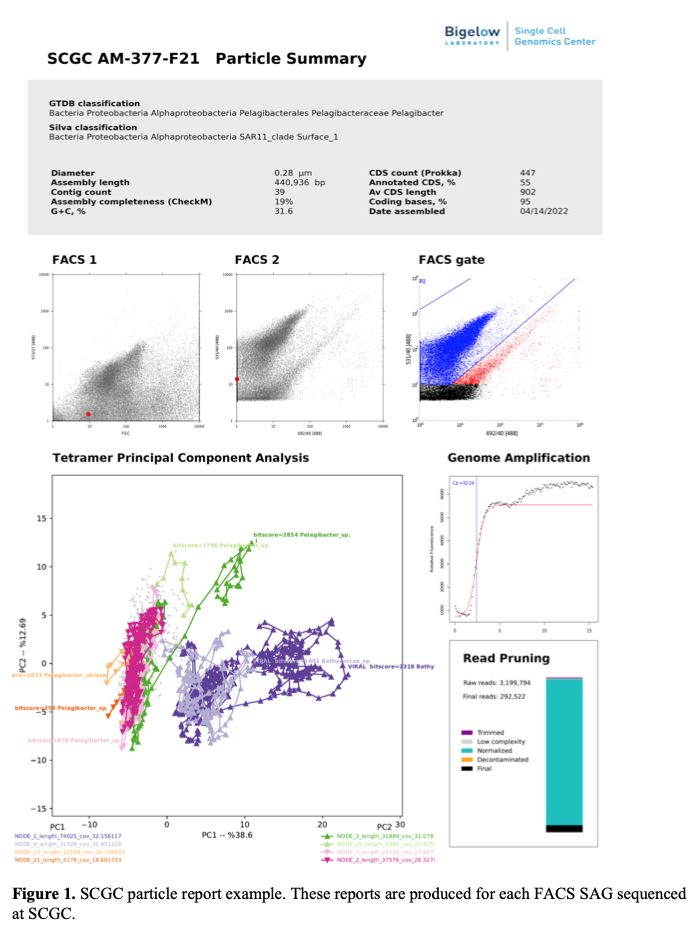
Single amplified genomes (SAGs) are products of whole genome amplification reactions performed on individual cells or other DNA-containing particles (1-3). To generate fluorescence-activated cell sorting (FACS)-based SAGs, cells are separated by FACS, lysed, and their DNA is amplified in 384-well microplates. SAGs are sequenced, de novo assembled, and annotated, with multiple steps in this workflow providing biologically meaningful data (Figure 1).
FACS SAG Generation WGA-X (S-201)
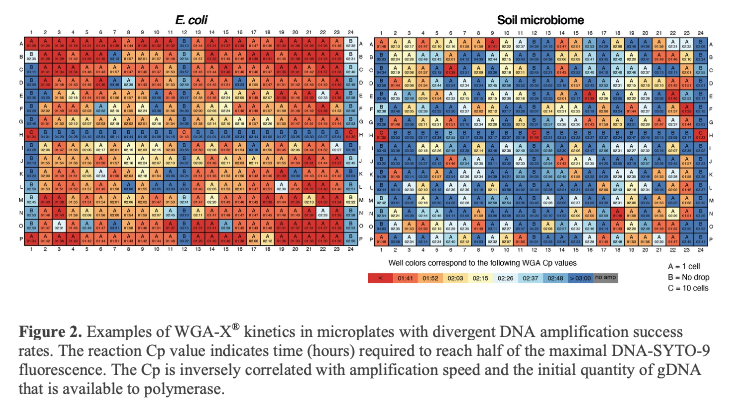
The S-201 service includes separation of individual cells or DNA-containing particles into wells of a 384-well plate by fluorescence-activated cell sorting (FACS), followed by cell lysis and genomic DNA amplification. Cells/particles are separated using an inFlux Mariner (BD), which can be finely tuned to select individual cells or particles based on a range of optical characteristics. Cells/particles may be selected based on the particle autofluorescence, the fluorescent DNA stain SYTO-9 (Thermo Fisher Scientific, provided by SCGC) or other stains and probes that are applied by an SCGC customer prior to shipping to SCGC. During sorting, multiple wells are used as negative and positive controls on each plate (Figure 2). Only one sample can be used per microplate.
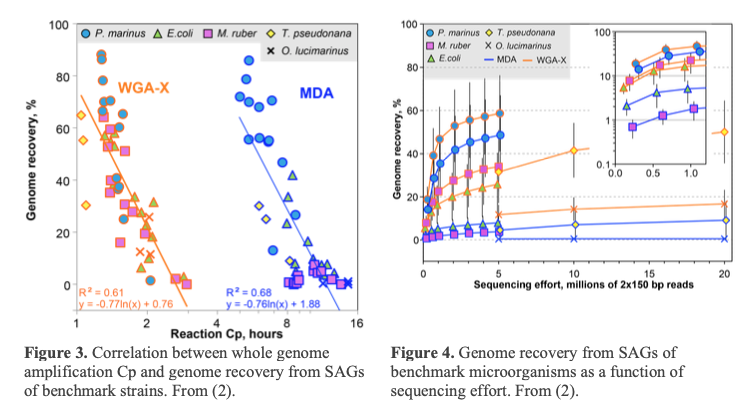
Cells/particles are lysed and their DNA is denatured by 2 freeze-thaw cycles and a subsequent KOH treatment (2). The S-201 service uses WGA-X® for genomic DNA amplification, a method developed by SCGC (2). Compared to the earlier versions of multiple displacement amplification technique (MDA) (4), WGA-X® improves average genome recovery from individual cells and viral particles, with most notable enhancements observed in SAGs with high G+C content (Figures 3 and 4). Please note that SAG generation success varies among samples (Figure 2) and depends on many factors, such as: a) prompt cryopreservation of intact cells and gDNA prior to cell sorting; b) successful discrimination of cells and viral particles from other particles during FACS; and c) successful single cell and viral particle lysis and DNA amplification.
Deliverables:
a) One 384-well microplate containing WGA-X® products of individual particles, 2 µL per well, usually averaging ~0.2 µg gDNA per well
b) Index FACS data files
c) WGA kinetics in each well
FACS SAG Generation WGA-Y (S-202)
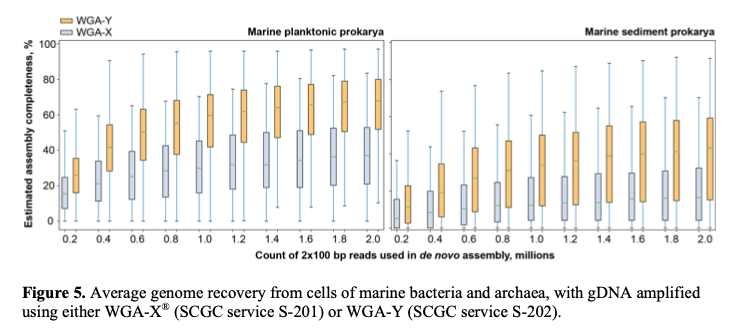
Service S-202 results in the same set of deliverables and involves the same requirements as S-201. The only difference of S-202 from S-201 is a modified genomic DNA amplification technique, WGA-Y, which enables a substantially improved average genome recovery from single cells (Figure 5). Although WGA-Y per se is more expensive than WGA-X®, it allows for shallower and cheaper sequencing of the resulting SAGs to produce similar or better genome recovery, which may reduce the overall project costs (Table 2). A manuscript describing the details of WGA-Y is in preparation.
Integrated Single Cell Respiration Measurement (S-203)
Service S-203 complements SAG generation (either S-201 or S-202) by measuring oxygen respiration rates of individual microbial cells (see reference 13). In order for SCGC to perform this service, customers must provide samples that have been pre-labeled with the RedoxSensor Green probe (Thermo Fisher Scientific), following protocol linked here. Please note that this method has not been fully validated for anaerobic respiratory processes.
Deliverables: Estimates of O2 respiration rates of the same individual cells that were used in SAG generation.
FACS SAG WGS 1.8 Billion Reads (S-211)
This service is best suited for sequencing large batches of SAGs to maximize genome recovery. DNA sequencing is performed using an Illumina NextSeq 2000 instrument and P4 reagents in 2×100 bp mode, yielding ~1.8 billion paired-end reads. The reads are demultiplexed with bcl2fastq (Illumina), trimmed with Trimmomatic (5), cleaned from low-complexity and reagent-contaminant reads, normalized with kmernorm, and assembled with SPAdes (6). This is followed by quality trimming and removal of contigs shorter than 2,000 bp, as previously described (2). Functional annotation is performed using Prokka (7), complemented by a custom protein annotation database compiled from Swiss-Prot (8) entries for Archaea and Bacteria. SAG taxonomic assignments are based on GTDB-Tk (14) and Silva (15). Estimates of genome assembly completeness and potential contamination are obtained with checkM (9). A complementary search for contaminant sequences is performed using tetramer principal component analysis (10). Please note that our current annotation tools are designed for bacterial and archaeal genomes and are not suitable for eukaryotes.
A total of up to 384 SAGs can be selected for sequencing from up to six 384-well SAG plates generated using SCGC services S-201 or S-202. SAG selection can be done by the customer or by SCGC, e.g., at random or based on the WGA-X® Cp values, which correlate with SAG’s potential for high genome recovery (Figure 3). The average SAG sequencing depth depends on the number of SAGs selected by a customer for sequencing, which can be guided by our prior results with benchmark cultures (Figure 4). Some SAGs may yield low read counts due to physical, biological, or biochemical factors within individual cells or samples, or to liquid-handling inconsistencies. Due to the high-throughput, low-cost nature of SCGC WGS services, they do not include free re-sequencing of such SAGs, unless the number of affected SAGs exceeds 10% of all sequenced SAGs.
Deliverables:
a) raw sequence reads
b) de novoSAG assemblies
c) functional annotation of bacterial and archaeal SAGs
d) SAG taxonomic assignments
e) general genome properties, such as GC content and coding density
f) files to assist manual QC: outputs of tetramer PCA, BLASTn and checkM
g) Single Particle Reports. These reports bring to your fingertips all key pieces of information about individual cells and other particles analyzed by SCGC, see example in Figure 1.
FACS SAG WGS 0.4 Billion Reads (S-212)
This service is similar to S-211; the only differences are: a) use of P2 sequencing reagents; b) lower cost; and c) the total read count is ~0.4 billion, i.e., ~3x lower than with service S-211. Service S-212 is best suited for full-depth sequencing of up to 384 prokaryotic SAGs produced by S-202 or up to 100 prokaryotic SAGs produced by S-201.
FACS SAG WGS 0.1 Billion Reads (S-213)
This service is similar to S-211; the only differences are: a) use of P1 sequencing reagents; b) lower cost; and c) the total read count is ~0.1 billion, i.e.; ~11x lower than with service S-211. Service S-213 is best suited for full-depth sequencing of a small number of SAGs or for low-coverage sequencing of up to 384 SAGs.
FACS SAG WGS Add-on 1.8 Billion Reads (S-221)
This service includes deeper sequencing, de novo assembly, functional annotation, and QC of SAGs for which compatible Illumina libraries have already been produced by a single service S-211, S-212, or S-213, making the same types of deliverables. The total number of new, 2×100 bp reads produced by this service is ~1.8 billion. More reads per SAG usually result in better genome recovery, but this relationship is not linear (Figure 4). Per the customer’s request, SCGC can generate lists of FACS SAGs that meet S-221 eligibility criteria and are either randomized or prioritize SAGs with the lowest WGA Cp values – indicators of the potential for high genome recovery (Figure 3). Please note that, due to potential liquid handling inconsistencies, some FACS SAGs may receive unexpectedly low numbers of sequence reads. Due to its high-throughput, low-cost nature, service S-221 does not include free re-sequencing of such SAGs, unless the number of affected SAGs exceeds 10% of all sequenced SAGs.
FACS SAG WGS Add-on 0.4 Billion Reads (S-222)
This service is similar to S-221. The only differences are: a) use of P2 sequencing reagents; b) lower cost; and c) the total read count is ~0.4 billion, i.e., ~3x lower than with service S-221.
FACS SAG WGS Add-on 0.1 Billion Reads (S-223)
This service is similar to S-221. The only differences are: a) use of P1 sequencing reagents; b) lower cost; and c) the total read count is ~0.1 billion, i.e., ~11x lower than with service S-221.
SCGC SERVICES: SEMI-PERMEABLE CAPSULE (SPC) SAGS NEW!
SAG generation and sequencing using SPCs is a revolutionary technology developed by Atrandi Biosciences and optimized for environmental microbiology by SCGC (16). Compared to FACS-based SAGs, SPC technology enables higher throughput and the sequencing of genomes from individual particles below the optical detection limits of FACS. Some limitations include the inability to analyze particles’ optical properties, to make sequencing selections based on those properties, and to perform add-on sequencing of individual SAGs.
SPC SAG Generation and Barcoding (S-311)
The customer’s sample is compartmentalized into semi-permeable microcapsules (SPCs) using microfluidic platforms, Onyx or Flux (Atrandi Biosciences). An alkaline treatment lyses the encapsulated particles, and the genomic DNA is amplified by WGA-Y using the same protocols as for FACS SAGs (service S-202), as reported in (16). The counts of total and DNA amplicon-containing SPCs are determined by a microscopic examination of SPCs after staining with the fluorescent DNA dye Syto-9. If the first encapsulation yields >1% SPCs containing amplification products (SAGs), the sample is diluted and reprocessed to reduce the fraction of SPCs containing more than one particle. The customer SAGs are mixed with E. coli SPC SAGs, serving as internal standards, at ~10:1 ratio. The SPC SAG combinatorial barcoding is performed using Atrandi Biosciences kits CKP-BARK8. A four-step, ligation-based combinatorial split-and-pool barcoding process generates a barcode diversity of 24^4 (331,776) unique variants. SCGC’s standard protocols are designed to maintain the frequency of genome assemblies derived from >1 DNA-containing particle at <1%.
Deliverables: 100 – 10,000 barcoded SPC SAGs from a single sample
SPC SAG Illumina Library Production (S-312)
A single Illumina library is generated from an aliquot of up to 2,500 barcoded SPC SAGs (products of S-311) using a unique index that can be multiplexed with up to 3 additional, compatible libraries.
Deliverables: One ready-to-sequence Illumina library of up to 2,500 SPC SAGs
SPC SAG WGS, 1.8 Billion Reads (S-321)
One or multiple libraries produced by S-312 and containing compatible indexes are pooled and sequenced on a NextSeq 2000 (Illumina) using a P4 200-cycle kit. The reads are demultiplexed with bcl2fastq (Illumina) and Pheniqs (18). The subsequent bioinformatics workflow is similar to the SCGC FACS SAG sequencing workflow. It includes trimming raw reads with Trimmomatic (5), removing low-complexity and reagent-contaminant reads, normalizing reads with kmernorm, assembling with SPAdes (6), quality trimming of contigs, and removing contigs shorter than 1,000 bp, as previously described (16). Functional annotation is performed using Prokka (7), complemented by a custom protein annotation database compiled from Swiss-Prot (8) entries for Archaea and Bacteria. SAG taxonomic assignments are based on GTDB-Tk (14) and Silva (15). Estimates of genome assembly completeness and potential contamination are obtained with checkM (9). A complementary search for contaminant sequences is performed using tetramer principal component analysis (10). The E. coli internal standard is detected and evaluated using QUAST (18). Please note that our current annotation tools are designed for bacterial and archaeal genomes and are not suitable for eukaryotes. Deliverables:
a) raw sequence reads
b) de novo SAG assemblies
c) functional annotation of bacterial and archaeal SAGs
d) SAG taxonomic assignments
e) general genome properties, such as GC content and coding density
f) files to assist manual QC: outputs of tetramer PCA, BLASTn and checkM
SPC SAG WGS, 0.4 Billion Reads (S-322)
This service is identical to S-321, except that sequencing is performed with a NextSeq 2000 P2 200-cycle kit, which typically produces about 0.4 billion paired-end reads.
SPC SAG WGS, 0.1 Billion Reads (S-323)
This service is identical to S-321, except that sequencing is performed with a NextSeq 2000 P1 200-cycle kit, which typically produces about 0.1 billion paired-end reads.
SCGC SERVICES: OTHER SERVICES
Biosafety Level 2 Handling (S-004)
Service S-004 complements FACS SAG and SPC SAG generation services when processing biosafety level 2 samples. This fee covers the costs of laboratory preparation and staff training for processing biohazardous materials.
FACS SAG Re-Arraying (S-005)
FACS SAGs are transferred from 384-well plates to 96-well plates. SCGC customer defines transfer volumes, source wells, and destination wells. Prior to the transfer, the destination wells are pre-filled with 5-150 uL of deionized water or 1x TE buffer, as specified by the SCGC customer. Deliverables include re-arrayed SAGs in a 96-well plate.
Sample cryoprotectant glyTE (S-009)
For SCGC customers’ convenience, we offer the preparation and shipment of SCGC’s recommended sample cryoprotectant glyTE. Deliverables of this service include preparation and shipment of 10 mL of 10x concentrated glyTE. Please note that SCGC’s recommended procedures for sample cryopreservation, including the recipe for the cryoprotectant glyTE, are posted on the SCGC website under Preparation and Shipment.
Consultation (S-011)
Basic support is included in the pricing of all SCGC services. When SCGC customers require more extensive help with study design and/or data interpretation (greater than two hours per project), we may request compensation for the associated labor costs via consultant fees. We may also request co-authorship on resulting publications for those SCGC scientists who are providing substantial, project-specific intellectual input.
Customized Services (S-020)
SCGC offers a wide range of customized services, including non-standard cell sorting, non-SAG sequencing, bioinformatics support, and method development. For more information, please contact SCGC manager Brian Thompson.
COST ESTIMATE: CASE STUDY
Let’s assume an external SCGC customer wants to: a) generate two 384-well plates of FACS-based SAGs using an enhanced gDNA amplification process WGA-Y; b) measure respiration rates of cells corresponding to those SAGs; c) perform whole genome sequencing of 192 SAGs from each of the two plates; and d) produce and sequence up to 5,000 SPC SAGs. Fees for this work are summarized in Table 2.
Deliverables:
- Two 384-well microplates of FACS SAG DNA with associated index FACS, single cell respiration, and gDNA amplification kinetics data
- Up to 384 annotated and QC-ed FACS SAG assemblies
- Up to 5,000 annotated and QC-ed SPC SAG assemblies
Table 2. Cost estimate for a hypothetical SCGC project.
| Service | Cat. # | Price per unit | # of units | Amount |
| FACS SAG Generation WGA-Y | S-202 | $3,700 | 2 | $7,400 |
| Integrated Single Cell Respiration Measurement | S-203 | $900 | 2 | $1,800 |
| FACS SAG WGS 0.4 Billion Reads | S-212 | $19,000 | 1 | $19,000 |
| SPC SAG Generation and Barcoding | S-311 | $15,000 | 1 | $15,000 |
| SPC SAG Illumina Library Production | S-312 | $2,000 | 2 | $4,000 |
| SPC SAG WGS, 0.4 Billion Reads | S-322 | $11,000 | 1 | $11,000 |
| Subtotal | $58,200 | |||
| Large Order Discount, 15% | ($8,730) | |||
| Total | $49,470 |
REFERENCES
- Stepanauskas R. 2012. Single cell genomics: An individual look at microbes. Current Opinion in Microbiology 15:613-620.
- Stepanauskas R, Fergusson EA, Brown J, Poulton NJ, Tupper B, Labonté JM, Becraft ED, Brown JM, Pachiadaki MG, Povilaitis T, Thompson BP, Mascena CJ, Bellows WK, Lubys A. 2017. Improved genome recovery and integrated cell-size analyses of individual uncultured microbial cells and viral particles. Nat Commun 8:84.
- Stepanauskas R, Sieracki ME. 2007. Matching phylogeny and metabolism in the uncultured marine bacteria, one cell at a time. Proceedings of the National Academy of Sciences of the United States of America 104:9052-9057.
- Dean FB, Hosono S, Fang LH, Wu XH, Faruqi AF, Bray-Ward P, Sun ZY, Zong QL, Du YF, Du J, Driscoll M, Song WM, Kingsmore SF, Egholm M, Lasken RS. 2002. Comprehensive human genome amplification using multiple displacement amplification. Proceedings Of The National Academy Of Sciences Of The United States Of America 99:5261-5266.
- Bolger AM, Lohse M, Usadel B. 2014. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30:2114-2120.
- Nurk S, Bankevich A, Antipov D, et al. 2013. Assembling single-cell genomes and mini-metagenomes from chimeric MDA products. Journal of Computational Biology 20:714-737.
- Seemann T. 2014. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068-2069.
- Bateman A, Martin MJ, O’Donovan C, et al. 2017. UniProt: The universal protein knowledgebase. Nucleic Acids Research 45:D158-D169.
- Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW. 2015. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. PeerJ PrePrints 2:e1346.
- Woyke T, Xie G, Copeland A, González JM, Han C, Kiss H, Saw JH, Senin P, Yang C, Chatterji S, Cheng JF, Eisen JA, Sieracki ME, Stepanauskas R. 2009. Assembling the marine metagenome, one cell at a time. PLoS ONE 4.
- Mitra A, Skrzypczak M, Ginalski K, Rowicka M. 2015. Strategies for achieving high sequencing accuracy for low diversity samples and avoiding sample bleeding using Illumina platform. PLoS ONE 10.
- Sinha R, Stanley G, Gulati GS, Ezran C, Travaglini KJ, Wei E, Chan CKF, Nabhan AN, Su T, Morganti RM, Conley SD, Chaib H, Red-Horse K, Longaker MT, Snyder MP, Krasnow MA, Weissman IL. 2017. Index switching causes “spreading-of-signal” among multiplexed samples in Illumina HiSeq 4000 DNA sequencing. bioRxiv:125724.
- Munson-McGee JH, Lindsay MR, Sintes E, Brown JM, D’Angelo T, Brown J, Lubelczyk LC, Tomko P, Emerson D, Orcutt BN, Poulton NJ, Herndl GJ, Stepanauskas R (2022). Decoupling of respiration rates and abundance in marine prokaryoplankton. Nature 612: 764–770.
- Parks DH, Chuvochina M, Chaumeil PA, Rinke C, Mussig AJ, Hugenholtz P (2020) A complete domain-to-species taxonomy for Bacteria and Archaea. Nat Biotechnol 38:1079-1086.
- Quast C, Pruesse E, Gerken J, Peplies J, Yarza P, Yilmaz P, Schweer T, Glöckner FO (2012) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Research 41:D590-D596.
- Weinheimer AR, Brown JM, Thompson B, Leonaviciene G, Kiseliovas V, Jocys S, Munson-Mcgee J, Gavelis G, Mascena C, Mazutis L, Poulton NJ, Zilionis R, Stepanauskas R (2025) Single-particle genomics uncovers abundant non-canonical marine viruses from nanoliter volumes. Nature Microbiology.
- Galanti L, Shasha D, Gunsalus KC (2021) Pheniqs 2.0: accurate, high-performance Bayesian decoding and confidence estimation for combinatorial barcode indexing. BMC Bioinformatics 22:359.
- Gurevich A, Saveliev V, Vyahhi N, Tesler G (2013) QUAST: Quality assessment tool for genome assemblies. Bioinformatics 29:1072-1075