Third Microbial Single Cell Genomics Workshop June 14-18, 2015 Organizing committee: Paul Blainey, Broad Institute David Emerson, Bigelow Laboratory for[…]
Read moreSearch Results for:
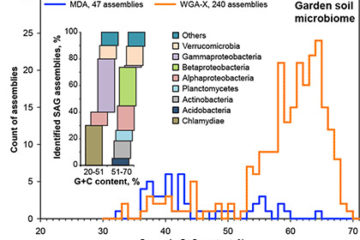
SCGC’s® WGA-X™ technology improves single cell genome recovery
Improved genome recovery and integrated cell-size analyses of individual uncultured microbial cells and viral particles Microbial single-cell genomics can be[…]
Read moreAdvisory Board
SCGC Advisory Board members on June 11th, 2019 Dr. Charles Budinoff Charles Budinoff is the Genomics and Microbiome Sciences Group[…]
Read moreTeam
Dr. Ramunas Stepanauskas, Director Brian Thompson, Manager Dr. Julia Brown, Research Scientist Dr. Gregory Gavelis, Bioinformatician Ben Tupper, Programmer Corianna Mascena, Research Associate SCGC® PARTNER[…]
Read moreHome
FAQ
How do I acknowledge SCGC services in my scientific publications and presentations?
Please make sure to indicate Bigelow Laboratory Single Cell Genomics Center in Materials and Methods in all publications resulting from[…]
Read moreHow do I name single amplified genomes (SAGs) generated by SCGC?
Please use the following convention: Escherichia coli SCGC ZZ-999-A01 Pseudomonas sp. SCGC ZZ-999-A02 Actinobacteria bacterium SCGC ZZ-999-A03 Euryarchaeon SCGC ZZ-999-A04 The use of this[…]
Read moreCan I collaborate with SCGC scientists beyond the scope of per-fee services?
Yes. General academic practices apply when collaborating with SCGC scientists beyond the scope per-fee services, such as reliance on mutual interests[…]
Read moreWhat methods does SCGC use?
A detailed description of SCGC services is provided here. For further details, please see: Stepanauskas R, Fergusson EA, Brown J, Poulton[…]
Read more